Gene Ontology
Gene ontology is an initiative to create a universal vocabulary for the characteristics of genes and gene products [1]. It consists of the following aspects:
1. Cellular Component
2. Molecular Process
3. Biological Function
Using AmiGo 1.8 it is simple to determine a list of cellular, molecular, and biological processes associated with a protein.
1. Cellular Component
2. Molecular Process
3. Biological Function
Using AmiGo 1.8 it is simple to determine a list of cellular, molecular, and biological processes associated with a protein.
Cellular Components
Figure 1: The localization of HFE [2]
|
Molecular Functions
|
Biological Processes
|
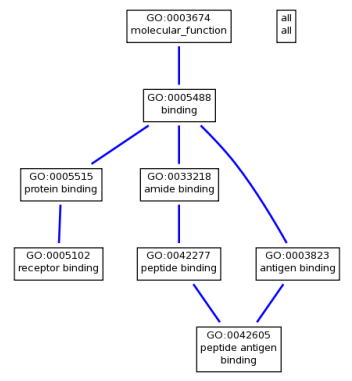
It is also possible to create trees that show the breakdown of a protein's functions, as seen below. To zoom in on the trees, either click the picture or download the file below the image.
Cellular Components
| gene_ontology_cellular_component.png | |
| File Size: | 162 kb |
| File Type: | png |
Molecular Functions
| gene_ontology_molecular_function.png | |
| File Size: | 23 kb |
| File Type: | png |
Biological Processes
| gene_ontology_biological_process.png | |
| File Size: | 692 kb |
| File Type: | png |
Analysis
The HFE protein localizes to the plasma membrane where it acts as a transmembrane protein. The cellular component gene ontologies for HFE make sense as the protein is transported to the membrane (hence the vesicles) and it largely localizes at the membrane (hence the large number of plasma membrane component gene ontologies). The molecular function ontologies are all related to a variety of forms of binding. This makes sense as the HFE protein interacts with TFR2 (Transferrin receptor 2) in a pathway that regulates iron absorption [3]. The biological functions are wide ranging, indicating that the protein participates in a variety of processes. However it is also clear that this protein is strongly involved in iron uptake and regulation.
References
1. Ashburner, M., Ball, C.A., Blake, J.A., Botstein, D., Butler, H., Cherry, J.M., Davis, A.P., Dolinski, K., Dwight, S.S., Eppig, J.T., Harris, M.A., Hill, D. P., Issel-Tarver, L., Kasarskis, A., Lewis, S., Matese, J.C., Richardson, J.E., Ringwald, M., Rubin, G.M., and Sherlock, G. (2000). Gene Ontology: tool for the
unification of biology. Nature Genetics, 25: 25-29.
2. Gruper. Y., Bar, J., Bacharach, E., and Ehrlich, R. (2005). Transferrin receptor co-localizes and interacts with the hemochromatosis factor (HFE) and the divalent metal transporter-1 (DMT1) in trophoblast cells. Journal of Cellular Physiology, 204: 901-912.
3. Goswami, T., Andrews, N.C. (2006). Hereditary hemochromatosis protein, HFE, interaction with transferrin receptor 2 suggests a molecular mechanism for mammalian iron sensing. Journal of Biological Chemistry, 281: 28494-28498. doi: 10.1074/jbc.C600197200
4. AmiGo! : http://amigo1.geneontology.org/cgi-bin/amigo/go.cgi
unification of biology. Nature Genetics, 25: 25-29.
2. Gruper. Y., Bar, J., Bacharach, E., and Ehrlich, R. (2005). Transferrin receptor co-localizes and interacts with the hemochromatosis factor (HFE) and the divalent metal transporter-1 (DMT1) in trophoblast cells. Journal of Cellular Physiology, 204: 901-912.
3. Goswami, T., Andrews, N.C. (2006). Hereditary hemochromatosis protein, HFE, interaction with transferrin receptor 2 suggests a molecular mechanism for mammalian iron sensing. Journal of Biological Chemistry, 281: 28494-28498. doi: 10.1074/jbc.C600197200
4. AmiGo! : http://amigo1.geneontology.org/cgi-bin/amigo/go.cgi