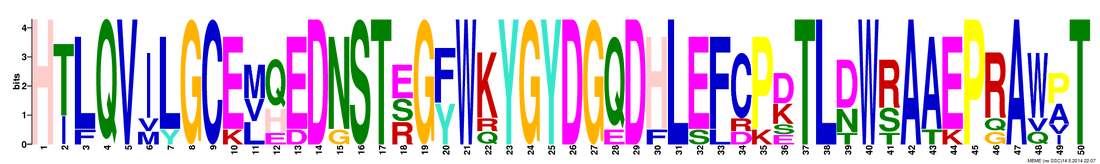Motifs
What are motifs?
In DNA, a sequence motif is a short reoccurring pattern of nucleotides that is thought to have conserved function [1]. These motifs are also found in the amino acid sequences in protein, and are thought to be important in the function of the proteins in which they are found [2]. As a result, motifs can be used to predict how a protein might function. In the case of hereditary hemochromatosis doing this work could give insight into how the mutations in HFE (H63D, S65C, and C282Y) play a role in disrupting the function of the HFE protein.
Data
In order to generate the following motifs I used a database called MEME. I input all of the protein sequences I found from my homologues. However, the program was only able to properly align the sequences from the vertebrates. Therefore, the motifs below are the alignment of the sequences from H. sapiens (Human), P. troglodytes (Chimpanzee), M. mulatta (Mouse), B. taurus (Cow), R. norvegicus (Rat), M. musculus (Mouse), and D. rerio (Zebrafish). A large letter within these motif images indicates complete conservation of an amino acid across species, while stacked letters indicate the variance in these amino acid positions.
Figure 1: Motif 1. This motif spans amino acids 116 to 165 in the human sequence.
Figure 2: Motif 2. This motif spans amino acids 61 to 110 in the human sequence. This means that it also contains two possible mutation sites in HFE: H63D (position 3 in the figure) and S65C (position 5 in the figure).
Figure 3: Motif 3. This motif spans amino acids 225 to 274 in the human sequence. It is just out of range of a possible mutation in HFE, C282Y, which occurs at position 282.
Analysis
By using this database I was able to glean quite a bit of information. First, the first motif does not span an area that contains one of the common HFE mutations. However, clinical studies have found that some hemochromatosis patients that are heterozygous for the C282Y mutation carry a novel mutation other than H63D or S65C [3]. Perhaps there are hemochromatosis patients that have a novel mutation in or near this motif. Given that this motif exists, it is likely that this portion of the protein is important for its overall function. Disrupting it could have similar effects as the other well known HFE mutations.
The second motif is quite interesting as it contains the mutation sites of H63D and S65C. The fact that these mutation sites are found in a motif tell me that there is a strong likelihood that these sites are indeed important for the overall function of HFE. This is interesting to note for my specific aims where I seek to identify how these mutation sites are important and why they can cause hemochromatosis when they do not disrupt the structure of the protein.
The third motif interests me because of what is missing. This motif spans amino acids 225 to 274 in the human sequence, which is just out of range for C282Y, the mutation most commonly associated with hereditary hemochromatosis. I find it very intriguing that the mutation that is the most likely to result in hemochromatosis is not found in any of the motifs. However, since this mutation is just outside the range for motif 3, I predict that C282Y's role in disrupting the structure of the protein also results in this motif falling in an improper position where it is no longer able to function.
The second motif is quite interesting as it contains the mutation sites of H63D and S65C. The fact that these mutation sites are found in a motif tell me that there is a strong likelihood that these sites are indeed important for the overall function of HFE. This is interesting to note for my specific aims where I seek to identify how these mutation sites are important and why they can cause hemochromatosis when they do not disrupt the structure of the protein.
The third motif interests me because of what is missing. This motif spans amino acids 225 to 274 in the human sequence, which is just out of range for C282Y, the mutation most commonly associated with hereditary hemochromatosis. I find it very intriguing that the mutation that is the most likely to result in hemochromatosis is not found in any of the motifs. However, since this mutation is just outside the range for motif 3, I predict that C282Y's role in disrupting the structure of the protein also results in this motif falling in an improper position where it is no longer able to function.
References
1. D'haeseleer, P. (2006). What are DNA sequence motifs? Nature Biotechnology, 24, 423-425. doi:10.1038/nbt0406-423
2. Hardin, J., Bertoni, G., and Kleinsmith, L. J. World of the Cell. 8th ed. New York: Pearson, 2012. Print.
3. Piperno, A., Arosio, C., Fossati, L., Vigano, M., Trombini, P., Vergani, A., and Mancia, G. (2000). Two novel nonsense mutations of HFE gene in five unrelated Italian patients with hemochromatosis. Gastroenterology, 119(2), 441-445.
2. Hardin, J., Bertoni, G., and Kleinsmith, L. J. World of the Cell. 8th ed. New York: Pearson, 2012. Print.
3. Piperno, A., Arosio, C., Fossati, L., Vigano, M., Trombini, P., Vergani, A., and Mancia, G. (2000). Two novel nonsense mutations of HFE gene in five unrelated Italian patients with hemochromatosis. Gastroenterology, 119(2), 441-445.