Conclusions and Future Directions
Discussion
Although it is well known that mutations in the HFE gene are responsible for hereditary hemochromatosis, there is still much that could be done to characterize the protein structurally. For example, I am interested in the H63D and S65C mutations, which are found in the MHC_1 domain of HFE.. Although these mutations need to be found in conjunction with a C282Y mutation [1], I am curious as to why these mutations contribute to hereditary hemochromatosis at all. I have not found any research to suggest that they disrupt the folding structure of HFE like the C282Y mutation. That lead me to wonder if these mutation sites and the domain in which they reside, MHC_1, are all that important for the overall function of HFE. My overall hypothesis was that these mutation sites and the MHC_1 domain are indeed important for HFE's role in iron uptake.
Specific Aim 1
My first aim was to determine how conserved the known mutation sites in human HFE are between vertebrates and invertebrates. I was interested in comparing these two groups because vertebrates have blood, so having iron regulation is more applicable to this disease than in invertebrates, which do not have blood. I would expect that the mutation sites in MHC_1, H63D and S65C, would be conserved across vertebrates. This would indicate that these sites and MHC_1 are indeed important for HFE's role in iron regulation. In order to approach this aim, I aligned the sequences of HFE homologues using Clustal Omega and examined the conservation of the human mutation sites across species.
Results
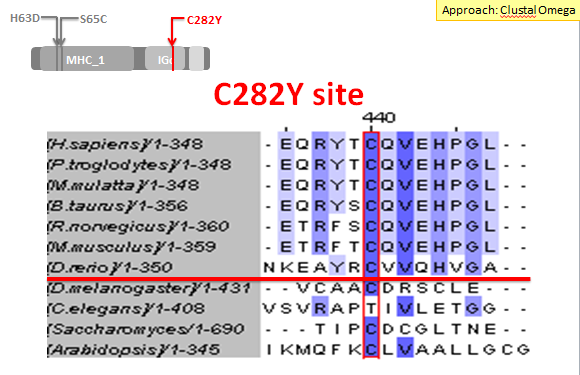
Although I was particularly focused on the H63D and S65C mutation sites, I also did a sequence alignment for the C282Y mutation site. There was significant conservation of this mutation across a multitude of species, not just vertebrates. This confirms that this mutation site is particularly important for HFE's function.
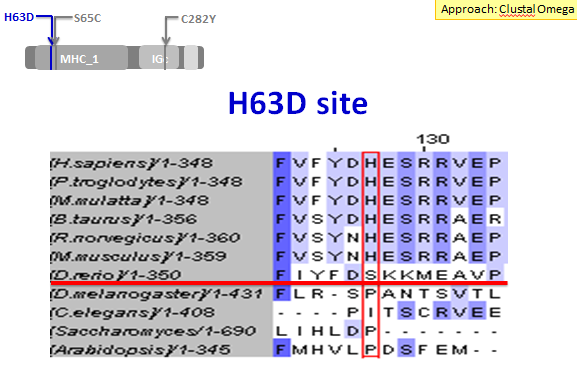
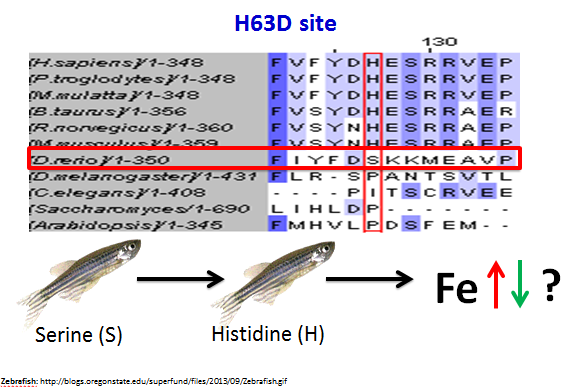
Next I examined the H63D mutation site. This site was not as highly conserved as the C282Y mutation site. In fact, it was not even entirely conserved across all of the vertebrates, as the zebrafish (D. rerio) did not have a histidine at this site like the other vertebrates. However, I feel that there is enough conservation across vertebrates to support the idea that this site and the MHC_1 domain are important for HFE's function.
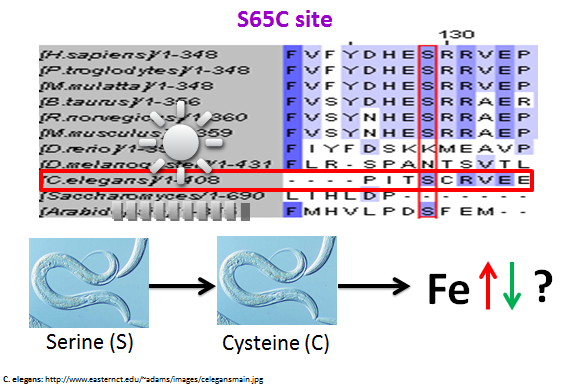
Finally, I examined the S65C mutation site. This site was particularly interesting because it was conserved across a variety of species, both vertebrates and invertebrates. Once again the zebrafish was not conserved at this site. However, C. elegans and Arabidopsis did have a serine at this site. Once again I feel that there is enough conservation to indicate that this site and the MHC_1 domain are important for the overall function of HFE. I also believe that it is possible to argue that the S65C site may be slightly more important for HFE than the H63D site due to the increased amount of conservation in the S65C site.
Future Directions
|
As a result of my research, I came up with two future directions of research that I think would be of interest to pursue. The first concerns the H63D site. The zebrafish has a serine at this site rather than a conserved histidine. Therefore, I think it would be interesting to mutate the sequence of the zebrafish homologue so that it contains a histidine at this position, rather than a serine, and examine the resulting iron levels to see if the homologue is still functional in the zebrafish. This would indicate if this amino acid at this site is indeed important for HFE function. It is possible that a serine simply performs a similar function to a histidine at this site in the zebrafish (Figure 4).
|
Another future direction that would be interesting to pursue is for the S65C site. Since there were invertebrates that had a conserved serine at this site, I thought it would be intriguing to induce a mutation in C. elegans where the serine at this site would be mutated into a cysteine, which is the exact mutation that occurs in human. Then I would look at the resulting iron levels to see if the function of the C. elegans protein homologue, DRAG-1, had been disrupted.
|
Specific Aim 2
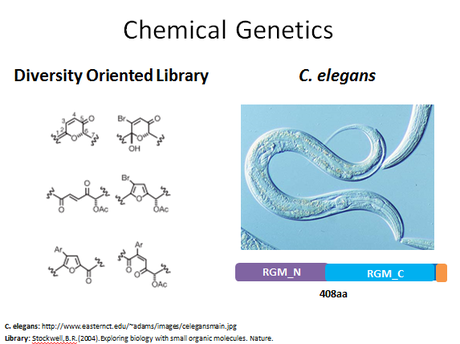 Figure 6: The chemical genetic approach for this aim
Figure 6: The chemical genetic approach for this aim
Since there are not any drug treatments for hereditary hemochromatosis, my second aim was focused on identifying potential drugs targets. Therefore, this aim is to identify drugs that can modify HFE protein function and lower iron levels. Since I have established that the MHC_1 domain is very likely important for HFE's role in iron uptake, I am focusing this aim on drugs that interact with the MHC_1 domain. My approach for this aim would be to use chemical genetics. I would use a diversity oriented library so that multiple types of organic molecules could be tested. My model organism would be C. elegans, as this is a very easy organism to work with and has a good homologue for human HFE. Although this organism does not have an MCH_1 domain, I feel that any drug targets identified in this study could be used in a future study with an organism that does have an MHC_1 domain.
|
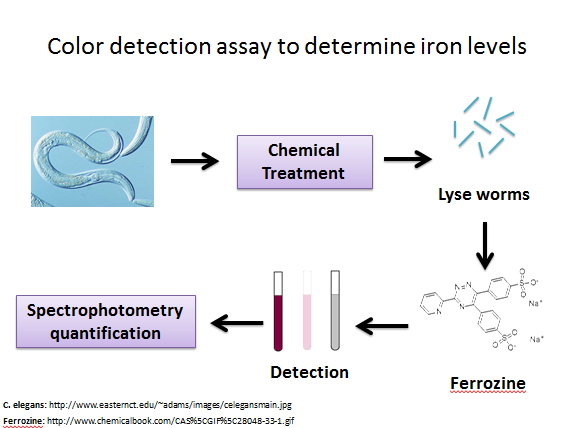
In order to perform a chemical genetics assay, I have formulated a color detection assay to detect iron levels in C. elegans. I have based this assay off of a ferrozine assay used on human brain astrocytes in culture [2]. In this detection process, C. elegans is first exposed to a small molecule from the library. Then the worms are lysed and exposed to a chemical called ferrozine. This chemical turns purple when it binds with iron, so a solution of lysed worm and ferrozine that is intensely purple indicates that there were high levels of iron in the worm (Figure 7). |
|
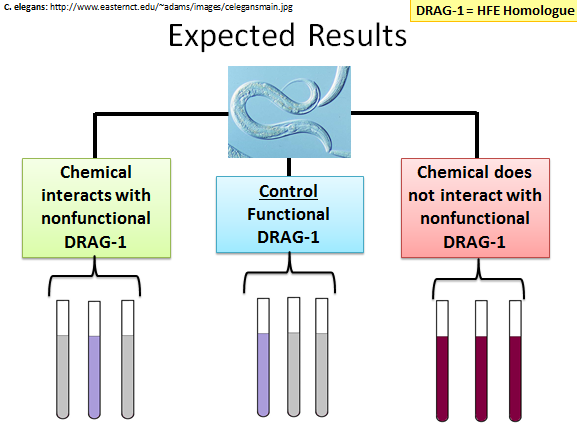
Figure 8 details the expected results depending upon what conditions are met. For this experiment the control is C. elegans with functional DRAG-1. I would expect that the iron levels in these worms would be very low, so there would not be a large amount of purple detected from the ferrozine assay. The experimental condition would be C. elegans with non-functional DRAG-1. If a chemical does interact with the mutated protein, then I would expect that it would restore DRAG-1's function and the ability of the worm to regulate iron would be restored. If a chemical does not interact with mutated DRAG-1, then iron regulation would not be restored and the ferrozine assay would turn intensely purple, indicating high levels of iron in the organism.
|
Future Directions
A future direction for this aim would be to take the chemicals that interacted with mutated DRAG-1 and test them in an organism that has an MHC_1 domain in their HFE homologue.
Conclusions
Specific aim 1: Due to its large amount of conservation across species, the C282Y mutation site is clearly important for the overall function of HFE. H63D and S65C, on the other hand, are not as conserved. However, I feel that there is still enough conservation to consider these mutation sites and the MHC_1 domain (which is where they are located) to be important for HFE function. Since S65D is more conserved across species, it could be argued that this site is more important for HFE than H63D.
Specific aim 2: A chemical that can interact with mutated DRAG-1 in C. elegans and restore the iron regulation of the organism could interact with the MHC_1 domain of human HFE to restore its function in regulating iron uptake. This chemical genetics experiment could be the starting point for identifying potential drugs for the treatment of hereditary hemochromatosis.
Specific aim 2: A chemical that can interact with mutated DRAG-1 in C. elegans and restore the iron regulation of the organism could interact with the MHC_1 domain of human HFE to restore its function in regulating iron uptake. This chemical genetics experiment could be the starting point for identifying potential drugs for the treatment of hereditary hemochromatosis.
| becky_reese_final_presentation_hfe.pdf | |
| File Size: | 1438 kb |
| File Type: | |
References
1. Piperno, A., Arosio, C., Fossati, L., Vigano, M., Trombini, P., Vergani, A., and Mancia, G. (2000). Two novel nonsense mutations of HFE gene in five unrelated Italian patients with hemochromatosis. Gastroenterology, 119(2), 441-445.
2. Riemer, J., Hoepken, H.H., Czerwinska, H., Robinson, S.R., and Dringen, R. (2004). Colorimetric ferrozine-based assay for the quantification of iron in cultured cells. Analytical Biochem, 331(2), 370-5
See Specific Aims page for more information
2. Riemer, J., Hoepken, H.H., Czerwinska, H., Robinson, S.R., and Dringen, R. (2004). Colorimetric ferrozine-based assay for the quantification of iron in cultured cells. Analytical Biochem, 331(2), 370-5
See Specific Aims page for more information