Gene Homology
What is homology?
|
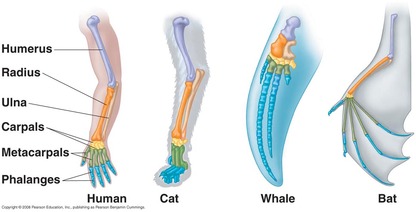
Figure 1: An example of homology among the bones of different species.
|
Homology is the idea that shared ancestry can be inferred based on the presence of similar structures between species [1]. The most commonly known image associated with homology is the one shown in Figure 1, where seemingly very different species share homologous bone structures, indicating that they likely share a common ancestor.
In addition to macro structures like bones, homology can be found in gene and protein sequences. This is done by comparing either genomic DNA sequences or protein amino acid sequences and determining how identical and similar they are. In the case of amino acid sequences, there may be instances where species do not share an identical amino acid but the corresponding amino acid may be similar enough (eg. they are both hydrophobic) and therefore have a similar function. These amino acids would be considered similar.
|
Methods
Possible homologs of the HFE gene were determined using Homologene, a database sponsored by the NCBI. Then the BLAST tool was used to search for sequences in other species that were homologous, or similar, to the human sequence. This tool is very effective for doing this because it aligns sequences from a library of genes and determines how many identical base pairs there are between the two sequences. This gives an estimate of how similar the human gene is to the homologous gene in another species.
Results and Analysis
Almost all of the homologues of the HFE protein are found in mammals. This makes sense, as HFE is involved in iron levels in the blood, so it will tend to only be found in animals that have some form of a cardiovascular system. When blasting for HFE homologous genes in primates, the chimpanzee had closest match to the human HFE gene with 99% identical matches. This was followed by the Rhesus macaque which had 93% identical matches. It is not surprising that primates would have such strong homology as they are considered human’s closest relative. The low E values indicate that this is highly statistically significant.
Interestingly, although there are many proteins that are homologous to the human HFE protein (see protein homology), as blasting was done on species that were more distantly related to humans it became more difficult to find homologous genes. Many of the species only had predicted homologous gene products. Other species, such as the rat (R. norvegicus) had matches to very vague portions of their chromosomes. In order to determine if post-transcriptional modifications were affecting the BLAST search results I compared mRNA from non-human species to the human sequence. The results, however, were inconclusive. This was the case regardless if the gene was considered “predicted” or not. The protein homology will likely provide more insight into the similarity across species than the gene homology in this case.
Interestingly, although there are many proteins that are homologous to the human HFE protein (see protein homology), as blasting was done on species that were more distantly related to humans it became more difficult to find homologous genes. Many of the species only had predicted homologous gene products. Other species, such as the rat (R. norvegicus) had matches to very vague portions of their chromosomes. In order to determine if post-transcriptional modifications were affecting the BLAST search results I compared mRNA from non-human species to the human sequence. The results, however, were inconclusive. This was the case regardless if the gene was considered “predicted” or not. The protein homology will likely provide more insight into the similarity across species than the gene homology in this case.
Homologue Reference Pages and Numbers
|
Human
H. sapiens Hemochromatosis (HFE) Accession Number: NG_008720.2 FASTA Rhesus macaque M. mulatta Mmul_051212 Accession Number: NC_007861.1 E value: 0.0 FASTA Identity: 93% |
Chimpanzee
P. troglodytes Hereditary Hemochromatosis HFE gene Accession Number: AF447807.1 E value: 0.0 FASTA Identity: 99% |
References
1. Understanding evolution: http://evolution.berkeley.edu/evolibrary/article/lines_04
2. Homologene: http://www.ncbi.nlm.nih.gov/homologene
3. NCBI: http://www.ncbi.nlm.nih.gov/
4. BLAST: http://blast.ncbi.nlm.nih.gov/Blast.cgi
2. Homologene: http://www.ncbi.nlm.nih.gov/homologene
3. NCBI: http://www.ncbi.nlm.nih.gov/
4. BLAST: http://blast.ncbi.nlm.nih.gov/Blast.cgi